The novel coronavirus doesn’t initially seem like a supercomputing problem. It’s a biological problem — a respiratory one, to be exact.
But the coronavirus poses modeling problems too. While the Trump administration has forecasted a pandemic death toll of between 100,000 and 240,000, that’s just a preliminary estimate. It’s “freaking hard” to forecast the death toll of the virus, FiveThirtyEight has noted, because the three key variables involved — the population susceptible, the infection rate and the fatality rate — remain open questions.
Take the infection rate: Thanks to limited testing, especially in the United States, we only have a count of the most acute coronavirus cases. How many people have had mild cases and recovered? How many had it without showing symptoms at all? It’s unclear.
The mortality rate, too, eludes us. Deaths seem easier to count than infections, but China may have undercounted its deaths, and the American numbers look similarly skewed.
The coronavirus’ novelty only compounds all this uncertainty. We’ve never seen it before, and scientists don’t precisely know how it infects us and spreads through our bodies. Much like its macro spread through the population, its smaller-scale spread through our respiratory systems is complex and full of unknowns.
“If we can understand how it works, then we can design things to break it.”
It’s essential to understand both, though. If we can model the spread of the coronavirus pandemic, we can plan appropriately at a policy level. If we can model the virus’ micro-level spread, meanwhile, that helps scientists develop vaccines and treatments.
“If we can understand how it works, then we can design things to break it,” Dr. Rommie Amaro, distinguished professor in theoretical and computational chemistry at the University of California–San Diego, told Built In.
So to learn more about the slippery virus, researchers have turned to supercomputers.
What Do a Virus Envelope and a Hypothetical City Have in Common?
Amaro leads a team modeling the coronavirus at the atomic level. Specifically, the team is building a model of the virus’ envelope — the spiky exterior you see in the ubiquitous Centers for Disease Control and Prevention rendering. This envelope is only 200 nanometers in diameter, Amaro estimates, but the modeling project is massive. The envelope alone contains about 200 million atoms.
Amaro isn’t just trying to create a digital statue of all these atoms, either. She wants her virus envelope to move.
“It’s four dimensional, really,” Amaro said, of the model. “It’s three dimensional, and then it’s also moving with respect to time.”
In Lemont, Illinois, researchers at Argonne National Laboratory also work on a model involving millions of moving parts. The moving parts aren’t atoms, though. They’re people, or “agents” — faceless, virtual figures who ping between millions of urban locations, ranging from four-person homes to 1,000-person workplaces.
They’re residents of CityCOVID, an agent-based model of Chicago that scientists can use to simulate different ways coronavirus could move through the city — and the different ways policy interventions could thwart it.
This complex creation, rooted in Chicago census data, works differently than the forecasting models that have dominated recent news. The Trump administration’s death forecasts, for instance, have relied on a model by Chris Murray, head of the Institute of Health Metrics and Evaluation.
That is a “largely statistical model,” noted Argonne’s social, behavioral and decision science group leader, Chick Macal.
It’s much quicker to mock up a statistical model than an agent-based one; Argonne’s has taken weeks to set in motion. However, statistical models also offer a “cruder way of modeling interventions,” Macal said. Creators have to guesstimate the impact of, say, closing schools, and manually tweak model parameters.
Not so with an agent-based model. To simulate school closures, researchers actually close the model’s digital schools, and change the agents’ hourly schedules accordingly. (Each agent has their own hourly schedule for weekdays, and a separate, fun schedule for weekends.)
Then they track how their agents fare. Each one has various possible coronavirus-related states: susceptible, infected, severely infected, hospitalized, and then either recovered or dead.
Amaro, for her part, hopes to ultimately track how models of human lung cells fare around her model coronavirus envelope. Her project’s underlying theory: if an atomically realistic model of a coronavirus envelope collides with a cluster of atomically realistic model lung cells, it will realistically infect those cells — and she and her team can observe the mechanics of the process up close.
Macal may be less interested in the close-up, but his and Amaro’s projects have more than coronavirus in common. Both use extremely granular models of things we know — the atomic composition of the virus; how people move around Chicago — to educate about the things we still don’t know. How does coronavirus infect us? How will it move through Chicago if the city does X, Y and Z?
Also, both take a lot of computing power.
How Researchers Use ‘Beefy Nodes’
For her model, Amaro relies on the Frontera supercomputer at the Texas Advanced Computing Center. This is the fifth-fastest supercomputer in the world, with almost half a million cores and peak processing power of about 23 petaflops. That means it can conduct 23 quadrillion (read: million billion) calculations per second.
It’s many thousands of times more powerful than an average laptop, which means researchers can use it, and machines like it, to complete projects in a few years that would have otherwise taken thousands of years. Leaning on parallel processing between many networked nodes — sometimes millions of them! — supercomputers accelerate sweeping, ambitious research projects.
These might involve mapping the human brain, simulating galaxies colliding, or, today, investigating coronavirus.
Amaro relies on supercomputers for her model’s raw material. It’s rooted in still images of the isolated coronavirus, taken by structural biologists through a process called cryo-electron microscopy. It captures the virus so close up that individual atoms appear as “hard balls,” Amaro said — and just developing these images requires supercomputing.
Amaro and her team then integrate the images with other data on coronavirus, drawn from mass spectrometry and other sources, ultimately translating two-dimensional representations into a four-dimensional model that can only run on a supercomputer.
It takes “hundreds of thousands of processors,” Amaro said.
That’s not just due to the millions of atoms, each located at shifting Cartesian coordinates; it’s the way those coordinates shift. The model’s motion is partially determined by an equation, Amaro explained, but there’s some “stochastic noise” in there too — in other words, some randomness in how the virus moves. It echoes the randomness of the natural world.
At Argonne, computational scientist Jonathan Ozik, too, compares the CityCOVID model to modeling nature.
“It’s kind of like a weather forecast,” he said.
However, CityCOVID models a slightly different kind of organic chaos — not randomness, but the inherent uncertainty in predicting the future with imperfect data. The Argonne team uses supercomputing to process and ultimately pare down that uncertainty.
The lab’s Theta supercomputer and Bebop HPC cluster can handle parameters that consist of ranges, Ozik explained.
Take R0, or “R naught.” Macal calls this “the single most important parameter” when it comes to modeling a virus: the average number of contacts a person with the virus infects.
Contact tracing — so, tracking how many of an infected person’s known contacts develop the virus themselves — has helped epidemiologists calculate the virus’ R0 in a “business as usual” world. However, estimates still vary, ranging from two to three.
In this context, that’s a huge gulf too; it’s the difference between cases doubling and tripling every few days.
To narrow it down, Argonne researchers can chop that two-to-three range into a few intervals, and see which one brings the model results closer to Chicago’s existing empirical data.
They’re not just trying to pinpoint R0, though. The CityCOVID model has about a dozen key parameters, Ozik said, all with their own ranges of possibilities. For instance — what’s the average length of time a person with COVID-19 is contagious but not symptomatic? What’s the probability per hour spent together that someone contagious will infect someone susceptible?
To narrow down all these ranges, Argonne researchers run multiple year-long CityCOVID simulations at once, with different permutations of parameters. Each year-long scenario, Ozik reported, takes about four of the Theta supercomputer’s “beefy nodes.”
“We have maybe 200 concurrent simulations running at a time,” he said.
But there’s no way to fully escape uncertainty. Even if Macal and Ozik find parameters that perfectly fit all the existing Chicago data, that data has its own inherent imperfections.
“There’s uncertainty in the data then gets kind of propagated or pushed forward to uncertainty ranges in the output,” Ozik said. “[It’s] what could happen given the best science knows so far.”
Amaro’s model, too, represents the best science knows so far. But both models are works in progress.

What’s Next for Pandemic Supercomputing?
Amaro, Macal and Ozik all agree that their models will take years to complete. So far, Amaro said, if we think of her model as a movie of the coronavirus envelope in action, “we’re still doing character development.”
To date, she and her team have a “beautiful” model of a red spike protein on the virus’ shell. This protein functions like a sensor, scanning its human host’s cells for an appropriate one to infect. Amaro’s team has already confirmed that this protein has “open” and “closed” positions — in the former position, it can infect, and in the latter, it’s harmless.
The factors that “close” it will grow clearer as the model develops.
“We’re going to learn something about what this spike protein is actually doing,” she said. “How is it interacting with antibodies?”
Argonne’s model, too, remains in the early stages. When Built In spoke with Macal and Ozik, they were finalizing their “business as usual” projections — in other words, their best guess as to what would have happened to Chicago if we had not intervened at all.
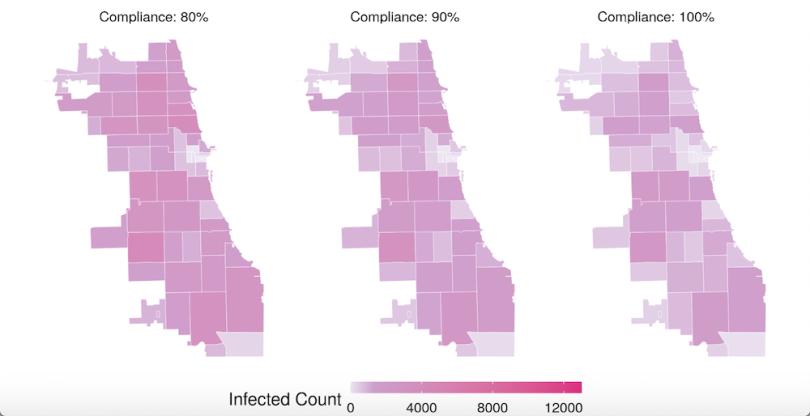
Ultimately, though, CityCOVID will calculate likely ranges of infection and mortality rates, given various possible interventions, and various possible levels of compliance.
They plan to share their findings with officials, and have already contacted the Chicago mayor’s office and the city’s department of public health.
One factor could throw a wrench in both modeling projects, though: mutations. Viruses evolve, and rapidly.
It’s unlikely mutations will radically alter how contagious or fatal the coronavirus is before we find a vaccine; scientists agree it’s “a slower-mutating virus,” Amaro said. In the longer term, though, mutations could change core aspects of how the virus passes between us and affects our bodies.
If that happens, Amaro would have to update her model, and Argonne’s team would have to update CityCOVID too.
“Our work can connect,” Amaro said.
Because ultimately, the macro and micro of this pandemic intersect — and not just because both require supercomputers.




